# Propagator Comparisons (IAS15 & DOP853)#
Last revised by Z. Ellis on 2026 MAR 30
Objectives#
This tutorial will demonstrate the usage of the two different integrators supported within Scarabaeus’ Propagator class, DOP853 and IAS15, across increasingly complex dynamics configurations. The configurations are:
Keplerian orbit
Keplerian orbit + third body perturbations
Keplerian orbit + third body perturbations + cannonball solar radiation pressure model
Keplerian orbit + third body perturbations + N-plate solar radiation pressure model
Keplerian orbit + third body perturbations + N-plate solar radiation pressure model + spherical harmonics gravity
The differences are highlighted by examining an inclined, elliptical geocentric orbit and noting where and why one integrator calculates different results than the other. Performance metrics like computational efficiency are also examined.
Imports and Set Up#
Here we’ll import the necessary libraries, define units and frames, and load a metakernel for SPICE time conversions.
[1]:
import scarabaeus as scb
import os, time
import pyrootutils
import numpy as np
import matplotlib.pyplot as plt
os.chdir(pyrootutils.setup_root(os.getcwd(), indicator = 'pyproject.toml', pythonpath = True))
## units, frames, kernels
kg, km, sec = scb.Units.get_units(['kg', 'km', 'sec'])
J2000 = scb.Frame('J2000')
scb.SpiceManager.load_kernel_from_mkfile(os.getcwd() + '/data/kernels/locked/locked_generic.tm')
Define Propagation Parameters#
Next, we’ll create an object to represent Earth, the body we’ll be orbiting in this tutorial, and its spherical harmonics information sh_info. We also need to create an orbiting spacecraft sc and an N-plate model for it.
We’ll also define the time interval we want to propagate over and a few orbital elements.
[2]:
## earth and spacecraft
earth = scb.CelestialBody.from_constants("EARTH")
sh_info = {'order' : 3,
'cs_file' : 'data/dynamic_setup/sph_coefficients/Earth_100.json',
'body' : scb.EARTH,
'norm_flag': True}
n_plate = scb.nPlateModel('data/dynamic_setup/nplate_coefficients/VandV_case5a_nPlate_config.json')
sc = scb.Spacecraft(name = 'ORCCA_SC',
spice_id = -1000,
tot_mass = scb.ArrayWUnits(2000, kg),
area = scb.ArrayWUnits(1e-06, km**2),
ref_coeff = scb.ArrayWUnits(1.5, None),
n_plate_model = n_plate,
attitudeMode = 'nadir_pointing_to_sun',
nadirPointingAxis = np.array([-1, 0, 0]))
## orbital parameters
t0 = scb.SpiceManager.cal2et('2026 JAN 01 00:00:00.000')
tf = scb.SpiceManager.cal2et('2026 JAN 01 10:00:00.000') # 10 hours
epochs = scb.EpochArray(np.arange(t0, tf, 500), 'TDB')
# semi major axis, ecccentricity, inclination, gravitational parameter
a, e, i, mu = 25000, 0.5, np.deg2rad(35), earth.grav_param.values
We want to examine how our integrators differ from one another, and the point that we’ll likely see the largest difference will be at periapsis where the spacecraft is moving fastest. In order to see this better, we want to examine a few different trajectories with initial conditions at different points in the orbit. We’ll create a function state_at_nu which will find the state corresponding to a given true anomaly nu so we can test these different initial true anomalies when we start
propagating.
[3]:
def state_at_nu(nu):
"""
Function to calculate state at given true anomaly nu.
"""
# perifocal position and velocity
h = np.sqrt(mu*a*(1 - e**2)) # angular momentum
r = h**2 / (mu*(1 + e*np.cos(nu)))
r0 = np.array([r*np.cos(nu), r*np.sin(nu), 0])
v0 = np.array([-(mu/h)*np.sin(nu), (mu/h) * (e + np.cos(nu)), 0])
# rotate to intertial intertial
peri_to_n = np.array([[1, 0, 0],
[0, np.cos(i), -np.sin(i)],
[0, np.sin(i), np.cos(i)]])
r0 = (peri_to_n @ r0.reshape((3, 1))).flatten()
v0 = (peri_to_n @ v0.reshape((3, 1))).flatten()
# convert to AWF and return
pos0 = scb.ArrayWFrame(r0, km , J2000)
vel0 = scb.ArrayWFrame(v0, km/sec, J2000)
return (pos0, vel0)
Set up Propagation (and Plotting)#
Since we want to look at a few different propagation configurations, we’ll define another function that can:
propagate any arbitrary dynamics configuration
plot the resulting propagation
We’ll do this under prop_and_plot_fm, which will first initialize a figure, then initialize four different initial conditions at different true anomalies using state_at_nu. For each true anomaly, it will then propagate the initial conditions using both the DOP853 and the IAS15 integrator and compare their differences as percents for each state component. We’ll also return the average percent differences so that we can look at how each set of dynamics changes the results.
[4]:
## plotting
def prop_and_plot_fm(model, title):
"""
Function to propagate the given dynamics using both DOP853 and IAS15
integrators. Plots the results and returns the average percent errors
for each state component.
Also returns average computation time for both integrators.
"""
# initialize plot
fig, axes = plt.subplot_mosaic([['o', 'o', '$X$', '$Y$', '$Z$'],
['o', 'o', '$\dot{X}$', '$\dot{Y}$', '$\dot{Z}$']],
per_subplot_kw = {'o' : {'projection' : '3d'}},
constrained_layout = True, figsize = (17, 9))
# examine four different initial true anomaly cases
nus = [-90, 0, 90, 180] # approaching peri, at peri, approaching apo, at apo
# save computation time for each propagator and differences between them
tot_avg_pos_err, tot_avg_vel_err, times, traj_errs = [], [], [], []
dop_time, ias_time = [], []
for j, nu in enumerate(nus):
# get state that corresponds to this case's true anomaly
pos0, vel0 = state_at_nu(np.rad2deg(nu))
state = [("position", 3, "estimated", "dynamic", sc, pos0),
("velocity", 3, "estimated", "dynamic", sc, vel0)]
state_vector = scb.StateArray(epoch = epochs[0],
origin = earth,
state = scb.StateDefinition.from_components(state))
# propagate with DOP853
dop_start = time.time()
prop_dop = scb.Propagator(primary_body = sc,
state_vector = state_vector,
tspan = epochs,
integrator = 'DOP853',
force_models = model,
propagate_STM = False)
prop_dop.propagate()
dop_time.append(time.time() - dop_start)
ys_dop = prop_dop.ys
# and IAS15
ias_start = time.time()
prop_ias = scb.Propagator(primary_body = sc,
state_vector = state_vector,
tspan = epochs,
integrator = 'IAS15',
force_models = model,
propagate_STM = False)
prop_ias.propagate()
ias_time.append(time.time() - ias_start)
ts_ias = prop_ias.times.times.values
ys_ias = prop_ias.ys
times.append(ts_ias)
## plot 3d orbit
# IAS15
if j == 0: # only need legend labels for one
axes['o'].plot(ys_ias[:, 0], ys_ias[:, 1], ys_ias[:, 2],
'b', label = 'IAS15')
axes['o'].plot(ys_ias[0, 0], ys_ias[0, 1], ys_ias[0, 2],
'go', label = 'IAS15 $t_0$')
axes['o'].plot(ys_ias[-1, 0], ys_ias[-1, 1], ys_ias[-1, 2],
'rs', label = 'IAS15 $t_f$')
else:
axes['o'].plot(ys_ias[:, 0], ys_ias[:, 1], ys_ias[:, 2],
linewidth = 1.4)
axes['o'].plot(ys_ias[0, 0], ys_ias[0, 1], ys_ias[0, 2], 'go')
axes['o'].plot(ys_ias[-1, 0], ys_ias[-1, 1], ys_ias[-1, 2], 'rs')
# note which true anomaly only for IAS15
match nu:
case -90: offset = (0, 1000, 0, 1000)
case 0 : offset = (1000, -1000, 1000, 1000)
case 90 : offset = (0, 1000, 0, 2000)
case 180: offset = (0, -2000, 0, 1000)
axes['o'].text(ys_ias[0, 0]+offset[0], ys_ias[0, 1], ys_ias[0, 2]+offset[1],
f'\u03bd$_0 =$ {nu}$^\circ$')
axes['o'].text(ys_ias[-1, 0]+offset[2], ys_ias[-1, 1], ys_ias[-1, 2]+offset[3],
f'\u03bd$_0 =$ {nu}$^\circ$')
# DOP853
if j == 0: # only need legend labels for one
axes['o'].plot(ys_dop[:, 0], ys_dop[:, 1], ys_dop[:, 2],
'k--', alpha = 0.6, label = 'DOP853')
axes['o'].plot(ys_dop[0, 0], ys_dop[0, 1], ys_dop[0, 2],
'm.', label = 'DOP853 $t_0$')
axes['o'].plot(ys_dop[-1, 0], ys_dop[-1, 1], ys_dop[-1, 2],
'b+', label = 'DOP853 $t_f$')
else:
axes['o'].plot(ys_dop[:, 0], ys_dop[:, 1], ys_dop[:, 2],
'--', linewidth = 0.9)
axes['o'].plot(ys_dop[0, 0], ys_dop[0, 1], ys_dop[0, 2], 'm.')
axes['o'].plot(ys_dop[-1, 0], ys_dop[-1, 1], ys_dop[-1, 2], 'b+')
# percent errors
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
traj_errs.append(err)
# calculate average percent errors and save for this initial true anomaly
avg_pos_err = np.mean(err[:, 0:3])
avg_vel_err = np.mean(err[:, 3:6])
tot_avg_pos_err.append(avg_pos_err), tot_avg_vel_err.append(avg_vel_err)
# primary body
axes['o'].plot(0, 0, 0, 'ko')
# formatting
axes['o'].set_title(f'IAS15 Avg Computation Time = {np.mean(ias_time):.3f} sec\n'
f'DOP853 Avg Computation Time = {np.mean(dop_time):.3f} sec')
axes['o'].legend(bbox_to_anchor = (0.5, -0.15), loc = 'lower center', ncols = 2)
axes['o'].set_aspect('equal'), axes['o'].view_init(elev = 33, azim = -60)
## plot percent errors for each initial true case
# plot values with smaller average error in front of ones with larger
zorder_inds = sorted(range(len(traj_errs)), key = lambda err: np.mean(err), reverse = True)
traj_errs = [traj_errs[j] for j in zorder_inds]
nus = [nus[j] for j in zorder_inds]
for t, traj_err, nu in zip(times, traj_errs, nus):
for j, ax in enumerate(axes):
if ax != 'o':
axes[ax].plot((t - t0)/3600, traj_err[:, j - 1], '-o', markersize = 3,
label = f'\u03bd$_0 =$ {nu}$^\circ$')
axes[ax].set_xlabel('Time [hrs]')
axes[ax].set_ylabel('IAS15 - DOP853 % Difference')
axes[ax].set_title(ax + f'\n Mean Difference: {np.mean(err[:, j - 1]):.4e} %')
axes[ax].grid(), axes[ax].legend()
# finalize plot and return average valeus
fig.suptitle(title)
plt.show()
return (np.mean(tot_avg_pos_err), np.mean(tot_avg_vel_err))
Dynamics Models and Analysis#
Now that we have a function that can propagate and plot any dynamics we give it, we need to define those dynamics. As a reminder, these are the five dynamics cases we will examine:
Keplerian orbit
Keplerian orbit + third body perturbations
Keplerian orbit + third body perturbations + cannonball solar radiation pressure model
Keplerian orbit + third body perturbations + N-plate solar radiation pressure model
Keplerian orbit + third body perturbations + N-plate solar radiation pressure model + spherical harmonics gravity
We’ll define each of these cases using ForceModelTranslation and then feed it to prop_and_plot_fm to compare the two propagators. We’re also saving the percent differences computed by prop_and_plot_fm to compare the two integrators for every force model later.
[5]:
## create and plot increasingly complex dynamics
# two body orbit
twobp = scb.ForceModelTranslation(primary_body = sc)
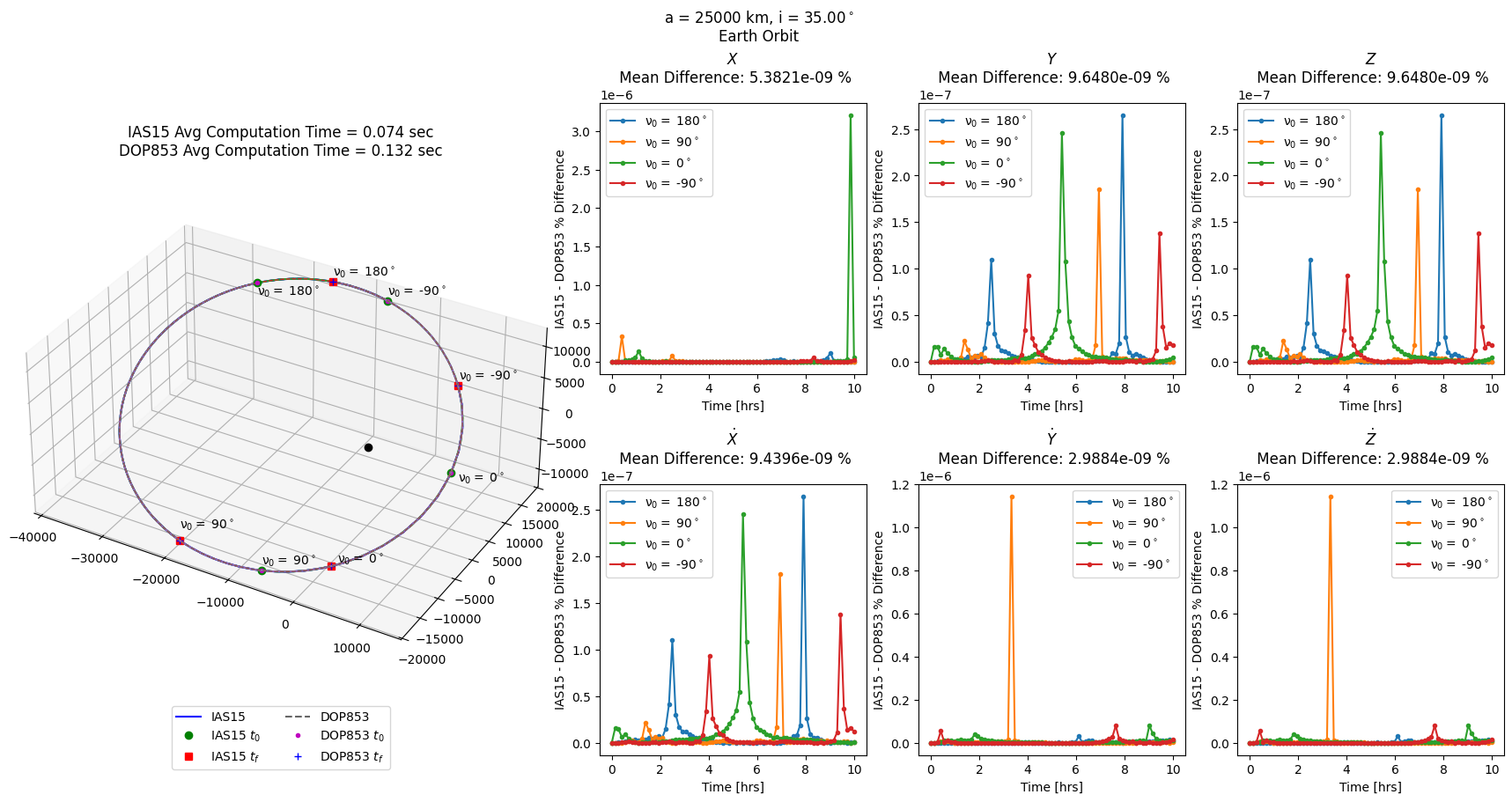
twobp_errs = prop_and_plot_fm(twobp, f'a = {a} km, i = {np.rad2deg(i):.2f}$^\circ$\n'
'Earth Orbit')
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
/var/folders/_q/_0gmy5p50pbf60yk0vzw402c0000gp/T/ipykernel_79006/3087986379.py:98: RuntimeWarning: invalid value encountered in divide
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
Propagation complete.
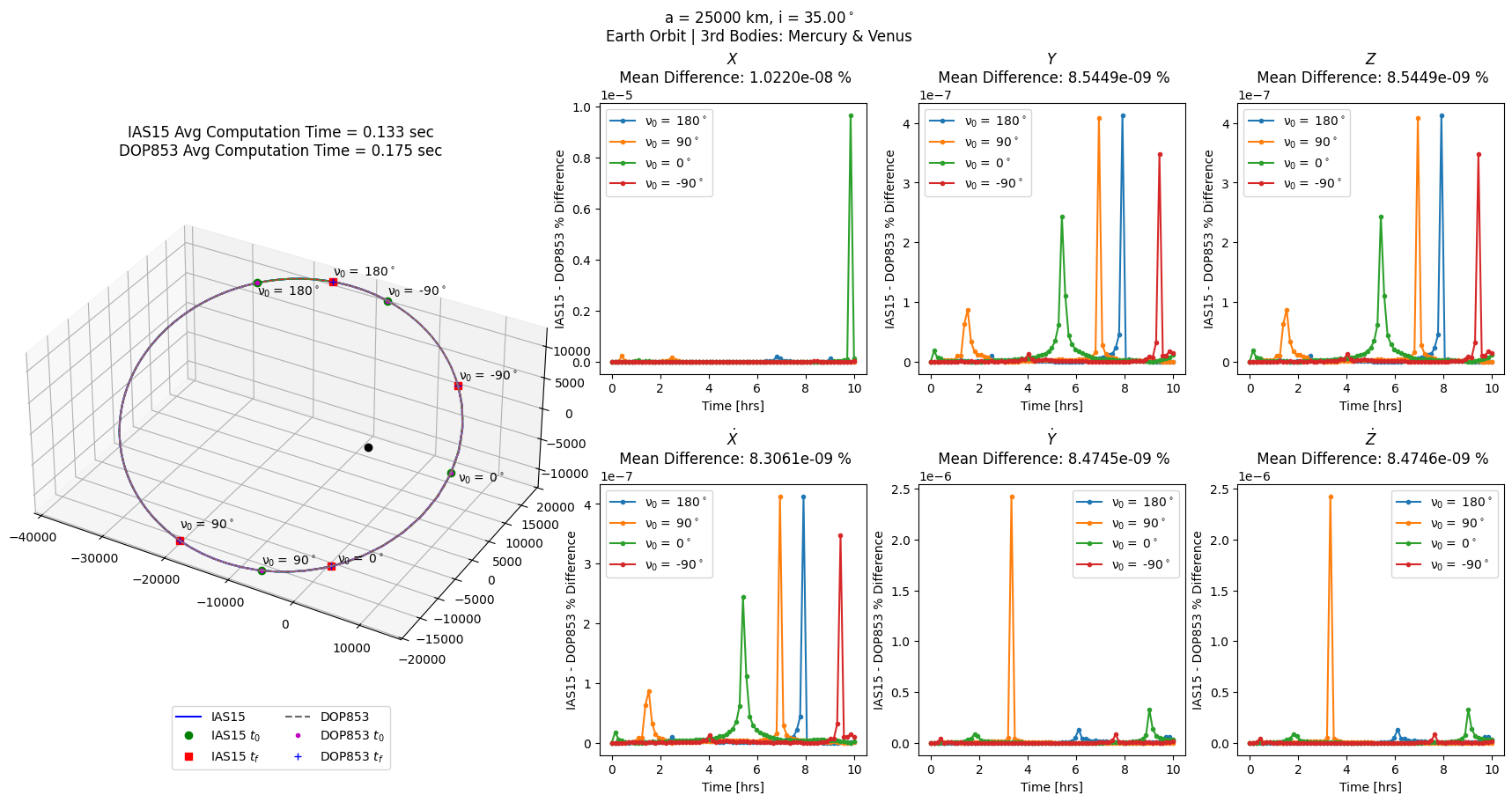
[6]:
# same as above with third bodies added
nBP = scb.ForceModelTranslation(primary_body = sc,
third_bodies = ['MERCURY', 'VENUS'])
nBP_errs = prop_and_plot_fm(nBP, f'a = {a} km, i = {np.rad2deg(i):.2f}$^\circ$\n'
'Earth Orbit | 3rd Bodies: Mercury & Venus')
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/var/folders/_q/_0gmy5p50pbf60yk0vzw402c0000gp/T/ipykernel_79006/3087986379.py:98: RuntimeWarning: invalid value encountered in divide
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
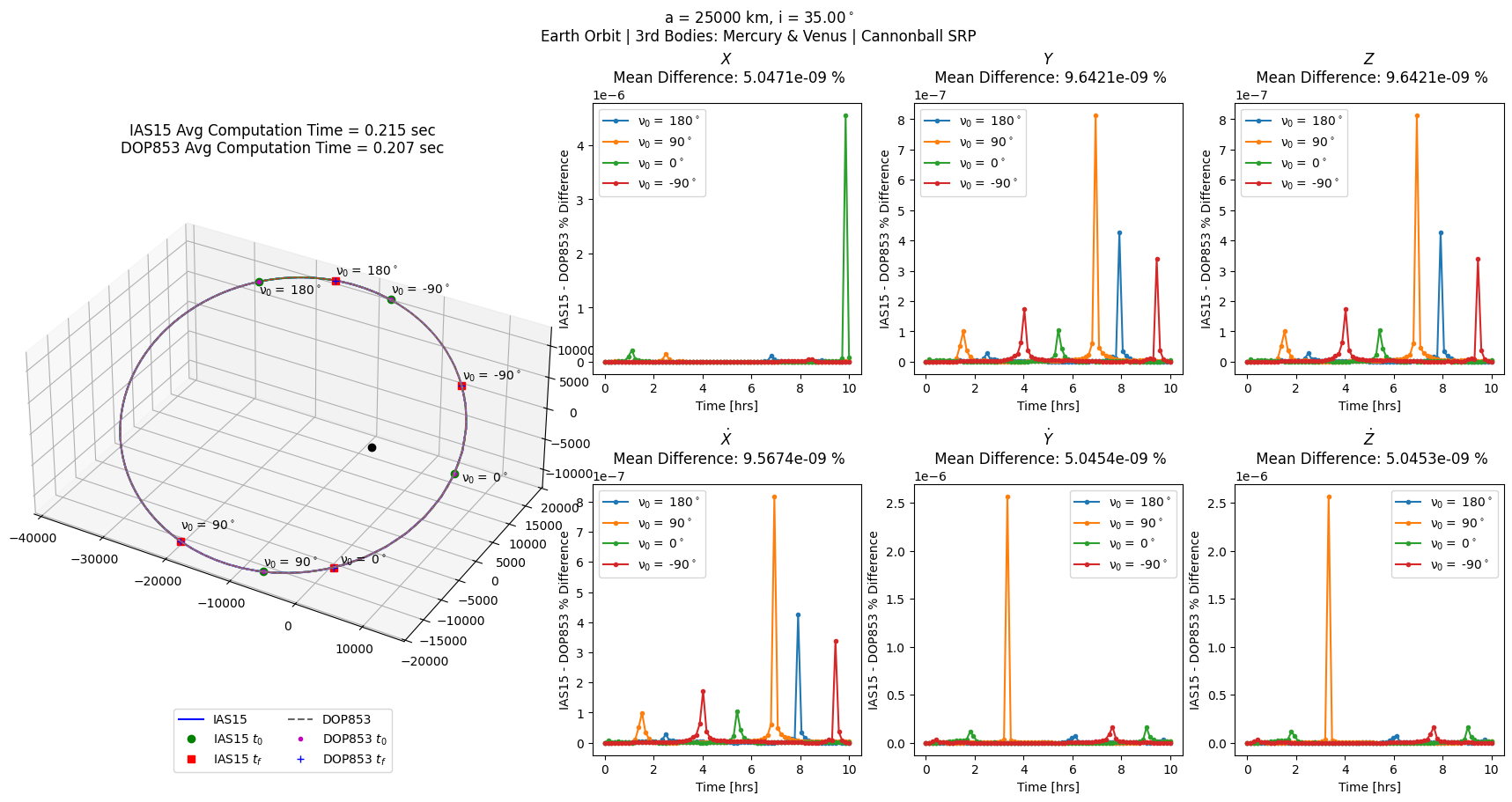
[7]:
# all above + cannonball SRP
cannon = scb.ForceModelTranslation(primary_body = sc,
third_bodies = ['MERCURY', 'VENUS'],
cannonball_SRP = True)
cannon_errs = prop_and_plot_fm(cannon, f'a = {a} km, i = {np.rad2deg(i):.2f}$^\circ$\n'
'Earth Orbit | 3rd Bodies: Mercury & Venus | Cannonball SRP')
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
/var/folders/_q/_0gmy5p50pbf60yk0vzw402c0000gp/T/ipykernel_79006/3087986379.py:98: RuntimeWarning: invalid value encountered in divide
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
=================== IAS15 integration complete. ==================
Propagation complete.
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
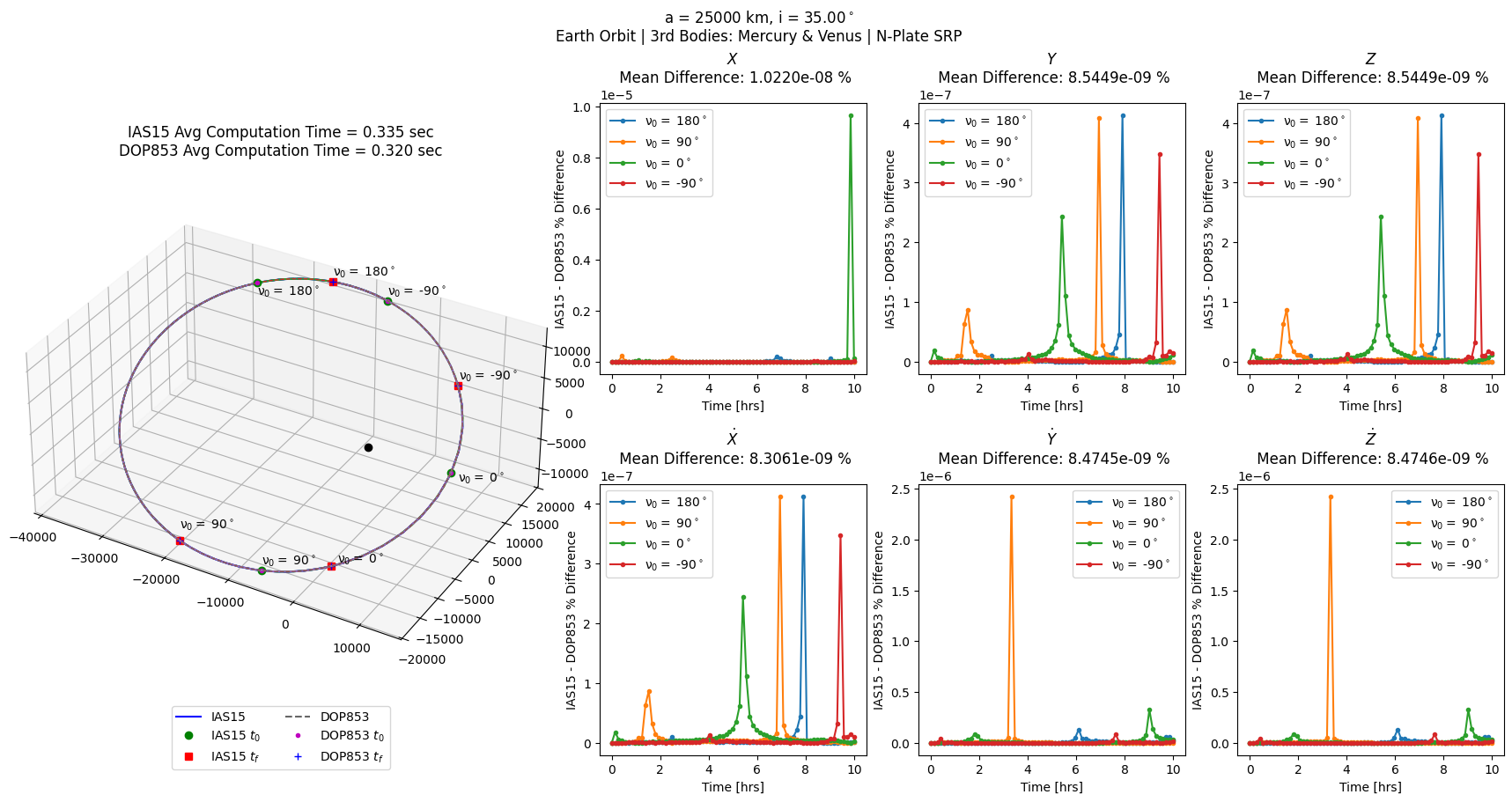
[8]:
# same as above with N-plate SRP instead of cannonball
npl = scb.ForceModelTranslation(primary_body = sc,
third_bodies = ['MERCURY', 'VENUS'],
nplate_SRP = True)
npl_errs = prop_and_plot_fm(npl, f'a = {a} km, i = {np.rad2deg(i):.2f}$^\circ$\n'
'Earth Orbit | 3rd Bodies: Mercury & Venus | N-Plate SRP')
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 0%| | 0.00/36000.00 s [00:00<?]/Users/zael5647/scarabaeus/src/scarabaeus/dynamics/ForceModelTranslation.py:568: RuntimeWarning: nadir_pointing_to_sun attitude mode not yet fully implemented for nPlateSRP. It currently produces incorrect partial derivatives.
model_acceleration = self.force_models[model].compute_acceleration(
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/var/folders/_q/_0gmy5p50pbf60yk0vzw402c0000gp/T/ipykernel_79006/3087986379.py:98: RuntimeWarning: invalid value encountered in divide
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
[9]:
# all above + spherical harmonics
sph = scb.ForceModelTranslation(primary_body = sc,
third_bodies = ['MERCURY', 'VENUS'],
nplate_SRP = True,
sph_harm = True,
sph_harm_order = sh_info['order'],
sph_harm_cs_file = sh_info['cs_file'],
sph_harm_body = sh_info['body'],
sph_harm_norm_flag = sh_info['norm_flag'])
sph_errs = prop_and_plot_fm(sph, f'a = {a} km, i = {np.rad2deg(i):.2f}$^\circ$\n'
'Earth Orbit | 3rd Bodies: Mercury & Venus | N-Plate SRP | 3rd Order SPH')
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/var/folders/_q/_0gmy5p50pbf60yk0vzw402c0000gp/T/ipykernel_79006/3087986379.py:98: RuntimeWarning: invalid value encountered in divide
err = np.nan_to_num(abs((ys_ias[:] - ys_dop[:, 0:6]) / ys_ias)*100, nan = 0)
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== IAS15 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
Integrating: 100%|███████████████████████████████████████████████| 36000.00/36000.00 s [00:00<00:00]
/Users/zael5647/scarabaeus/src/scarabaeus/utils/SCBPolynomial.py:80: RankWarning: Polyfit may be poorly conditioned
coeffs = np.polyfit(t_array, y_array, deg)
=================== DOP853 integration complete. ==================
Propagation complete.
================================================================================
Starting propagation...
================================================================================
=================== IAS15 integration complete. ==================
Propagation complete.
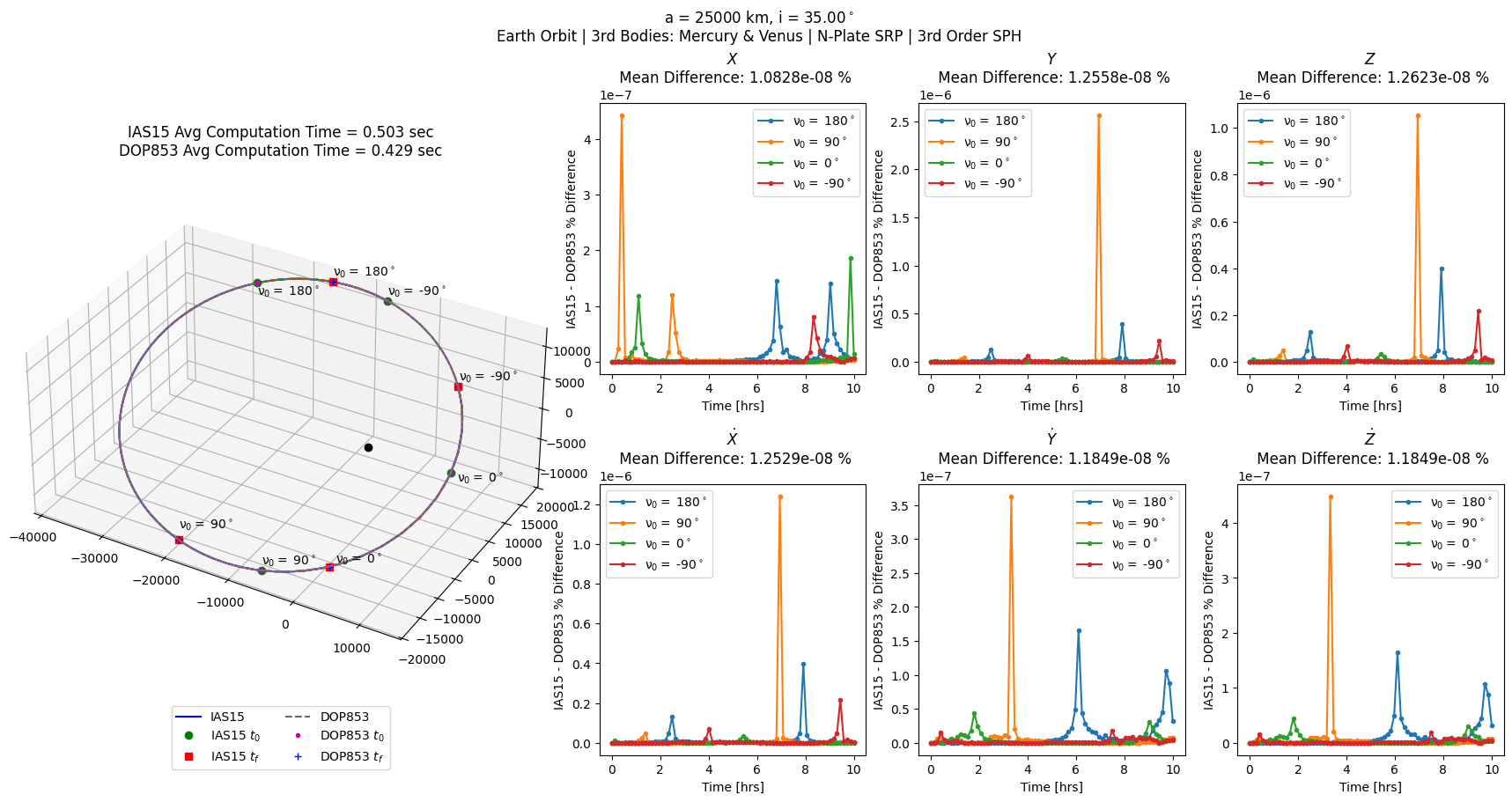
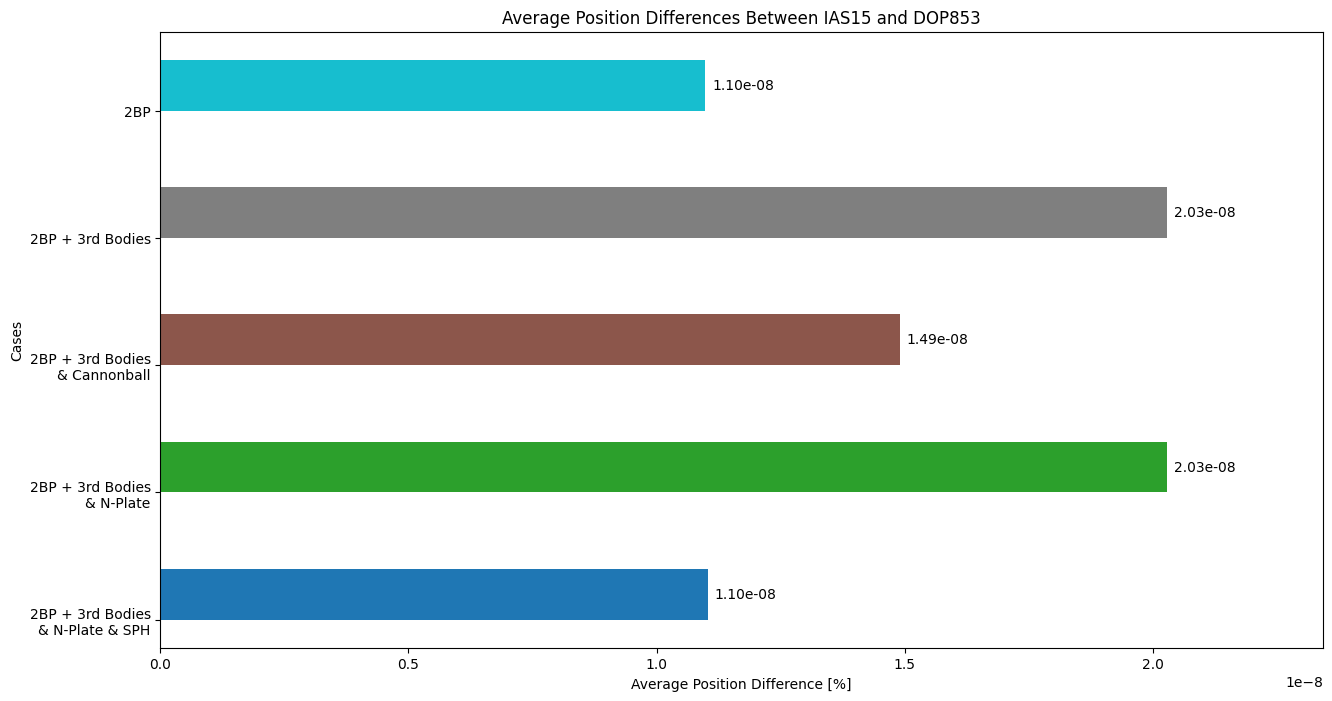
Now plot the average difference between DOP853 and IAS15 for position and velocity for all force model configurations.
[10]:
## plot average percent differences
# set up titles and colors
cases = ['2BP + 3rd Bodies\n& N-Plate & SPH', '2BP + 3rd Bodies\n& N-Plate',
'2BP + 3rd Bodies\n& Cannonball', '2BP + 3rd Bodies', '2BP']
ind, width = np.arange(len(cases)), 0.4
cmap = plt.get_cmap('tab10')
c = cmap(np.linspace(0, 1, len(cases)))
# position differences
fig_bar_pos, ax_bar_pos = plt.subplots(figsize = (15, 8))
# rust vs dop
bars = ax_bar_pos.barh(ind + width/2, [sph_errs[0], npl_errs[0], cannon_errs[0], nBP_errs[0], twobp_errs[0]], width, color = c)
ax_bar_pos.bar_label(bars, fmt='%.2e', padding=5)
ax_bar_pos.set(yticks = ind, yticklabels = cases)
ax_bar_pos.set_xlabel('Average Position Difference [%]'), ax_bar_pos.set_ylabel('Cases')
ax_bar_pos.set_xlim(0, 1.1*ax_bar_pos.get_xlim()[1]) # padding for bar labels
ax_bar_pos.set_title('Average Position Differences Between IAS15 and DOP853')
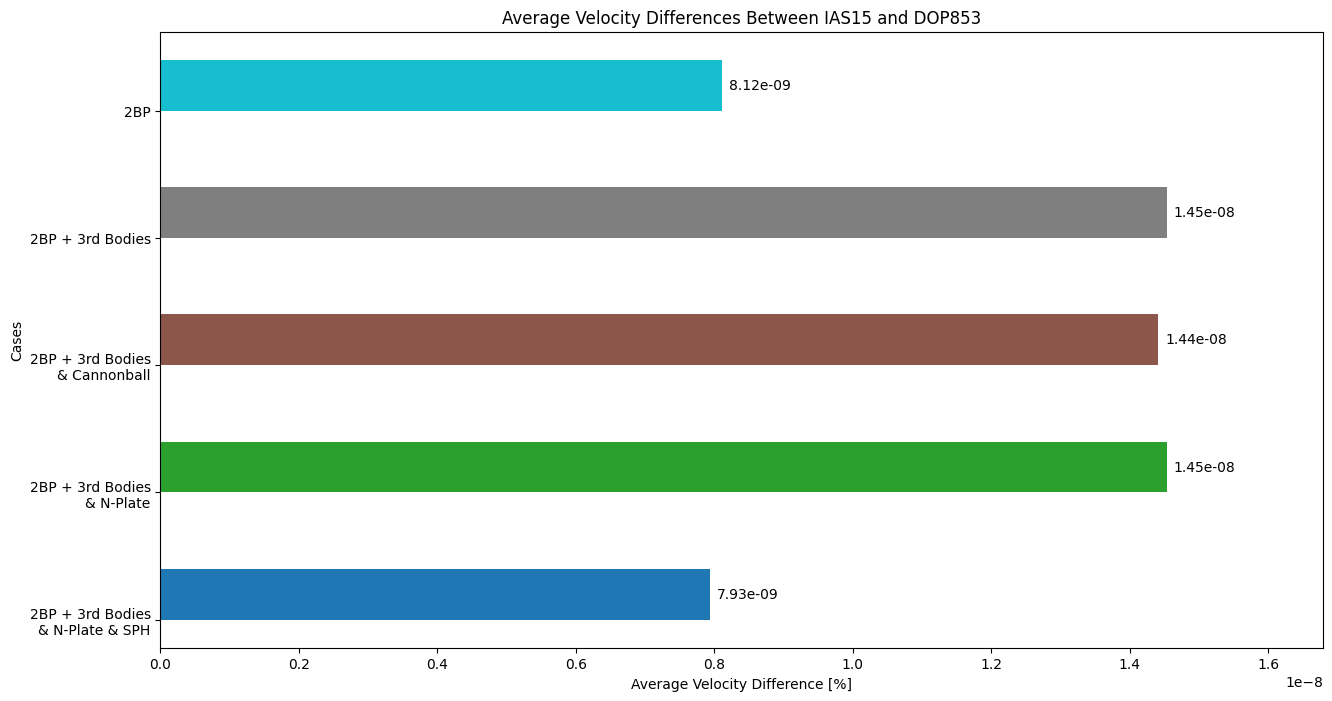
# velocity differences
fig_bar_vel, ax_bar_vel = plt.subplots(figsize = (15, 8))
# rust vs dop
bars = ax_bar_vel.barh(ind + width/2, [sph_errs[1], npl_errs[1], cannon_errs[1], nBP_errs[1], twobp_errs[1]], width, color = c)
ax_bar_vel.bar_label(bars, fmt='%.2e', padding=5)
ax_bar_vel.set(yticks = ind, yticklabels = cases)
ax_bar_vel.set_xlabel('Average Velocity Difference [%]'), ax_bar_vel.set_ylabel('Cases')
ax_bar_vel.set_xlim(0, 1.1*ax_bar_vel.get_xlim()[1]) # padding for bar labels
ax_bar_vel.set_title('Average Velocity Differences Between IAS15 and DOP853')
[10]:
Text(0.5, 1.0, 'Average Velocity Differences Between IAS15 and DOP853')
Conclusion#
FILL OUT
